Images capture critical moment of protein breakdown in diseases like Alzheimer’s: ScienceAlert
To perform a myriad of functions within the body’s cells, many types of hardworking protein slip in and out of dense droplets called condensates to Speed up biochemical reactions as it is required.
But that fluidity can break down in diseases — such as Alzheimer’s disease, Parkinson’s disease, and amyotrophic lateral sclerosis (ALS) — which are characterized by tough assemblages of clumpy proteins that form in nerve cells.
Now, a team of researchers has developed a new method for imaging the moment when proteins known to aggregate in neurodegenerative diseases begin to clump together.
Forming characteristic structures such as clumps, plaques, and interwoven fibrils, “proteins no longer exhibit rapid reversibility back to liquid form”. He explains Protein biophysicist Yi Xin, of the University of Sydney, who led the study.
“It is therefore important to monitor the dynamics of the condensate, as it directly affects disease states.”
Previous research has shown that proteins that aggregate in amyotrophic lateral sclerosis (ALS), a debilitating disease that affects motor function, are in a “supersaturated” state at very high concentrations. exceeds its typical solubility. In other words, these proteins teeter on the edge as soluble forms, and are liable to solidify if the cell becomes overwhelmed.
To get a closer look at the behaviors of such proteins, Shen and colleagues developed two new methods for closely monitoring the transition of a protein from its liquid to solid phase.
Their first test was a DNA/RNA-binding molecule called sarcoma protein (FUS), which aggregates in amyotrophic lateral sclerosis (ALS) and frontotemporal dementia.
When proteins such as FUS are concentrated as a gel in condensers, the dense, protein-rich phase is surrounded by a dilute phase depleted of particles. Bringing many proteins close together can cause the mixture to aggregate even further, irreversibly into solid clumps.
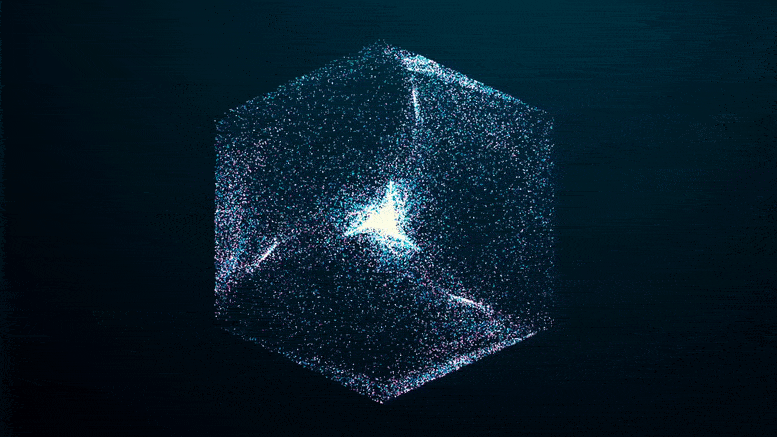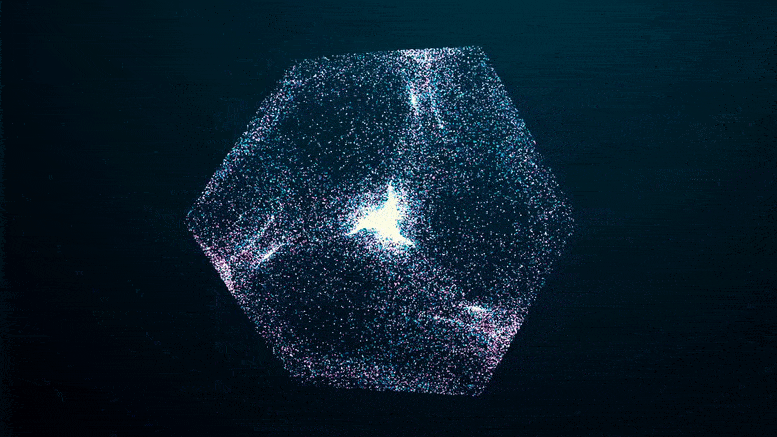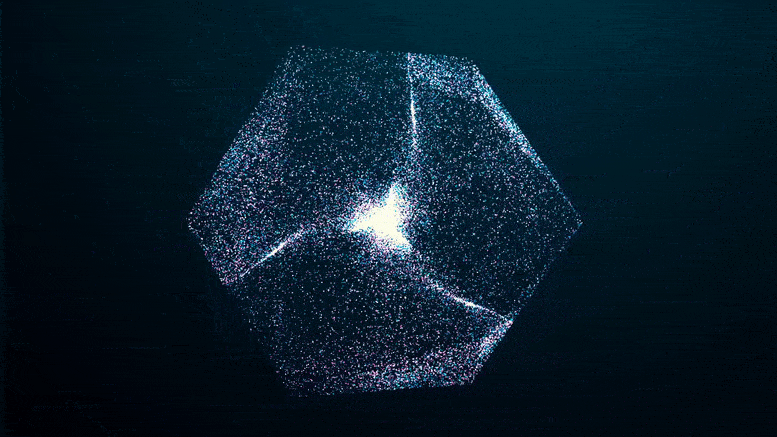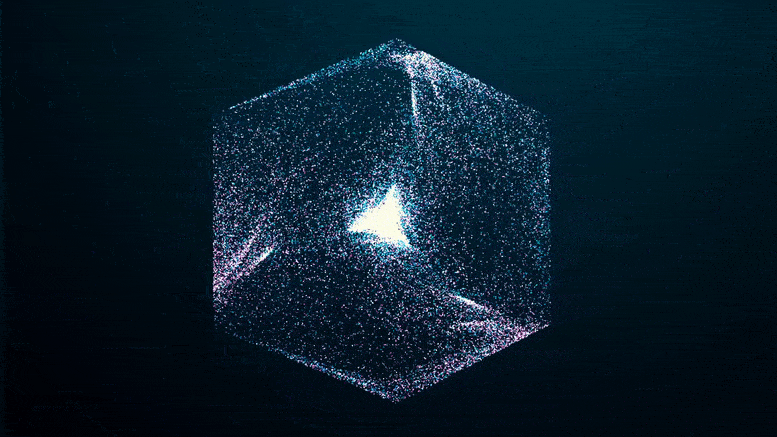
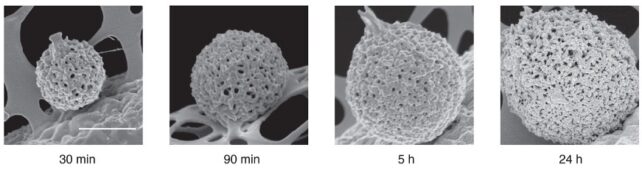
The researchers imaged solutions of FUS condensates as they formed over a 24-hour period, using two methods that collect light refracted by dense protein “spheres” and scatter it back. The patterns reflect the internal structure and density of the condensate, which hardened after five days.

“We use a fast camera to record long, bright field sequences of images at a high frame rate. This allows us to simultaneously explore fast dynamics (acquisition at high frame rates) and slow dynamics (through long-time acquisition)” Chen and colleagues. writing in their published paper.
Like an explosion in the night, you can see the green fluorescent proteins emerging from the blackness as they clump together, seeming to pull more and more proteins out of solution at the edges of the condensate until the whole clump appears to explode.
border frame=”0″ allow=”accelerometer; autoplay; clipboard writing; encrypted media; gyroscope; picture-in-picture; web sharing”>allowfullscreen
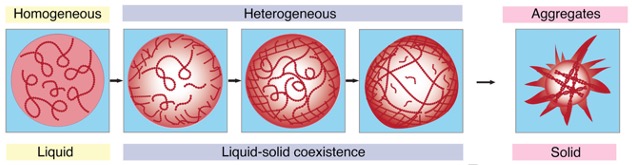
If you missed it in the time-lapse video above, watch it closely again: it shows that the transition from a liquid protein to a solid protein starts at the outer edge of the spherical condenser, which condenses and moves inward to the core until the entire droplet becomes a solid-like gel.
“This is a huge step forward in understanding how neurological diseases develop from a fundamental perspective.” He says Shane.
“We can now directly observe the transition of these important proteins from liquid to solid at the nanoscale – a millionth of a meter.” Add Daniele Vigolo, a biomedical engineer also at the University of Sydney.
While the experiments were conducted in solutions of proteins made in a lab, and there’s clearly a lot going on inside cells, the researchers say their findings provide new insights into the underlying physical process underlying neurodegenerative diseases.
If the imaging techniques can be replicated with other proteins, we might learn a lot about the strange ways in which these proteins interact.
The study was published in Banas.
Source link